Spreadsheets and Databases
Part 1: The four spreadsheet / database lessons I'm highlighting here are:
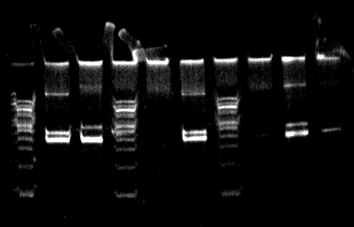
Graphing a standard curve: How to graph the results of a d1s80 PCR lab and calculate the size of your bands.
One of my favorite lessons in biotechnology was the d1s80 PCR lab because the allele frequencies are known for its 40+ alleles. Students ran PCR reactions on their own DNA in our class. One of the labs looked at D1s80, a specific site in our genome. There are a few reasons we do this lab. First, it offers more experience with doing PCR labs. Second, the difference in allele sizes is only 16 base pairs. Being so small, we had to use polyacrylamide vertical gel boxes just to be able to see the bands. We ran a 50 bp or 100 bp ladder so that we could take the measurement of the distance migrated for bands of known length. DNA moves logarithmically. When we know the size of the band, we can calculate the log of its size. This is when Excel comes in to play. I tried to do this with Google spreadsheets, but it could not handle either doing the log or the antilog. I just know that Google spreadsheets could not handle this level of analysis in 2009. We set up the log of the base pair length for the bands on the y axis and distance migrated in centimeters on the x axis. After we do a linear regression (line of best fit), Excel will give us an equation. We take the distance our bands migrated and plug it in to the equation for the value of x. Solving the equation gives us the log of the bp of our band. We take the anti-log and we know how big our bands are for d1S80. This is the main part of the lab that deals with spreadsheets. We would then go on and calculate the genotype frequency using data on the allele frequencies of all known band lengths, but I'm not sure elaborating on that falls into this lesson. You should be able to access the important parts of the lab from links I put in this explanation so if you wanted to create a hypothetical lab for your students, you could. The third reason for doing this lab, in case it is not obvious, is the exposure I could give students with not only how to use Excel, but let them get a taste for how powerful it can be. While doing a linear regression is not so amazing, I think it is for high school students. They'll probably have to do at least one in college. Having a clue what one is and how to do one before they get to college, can only prepare them for the competition they will face in their classes.
To see the student version of the curricula, go here: http://www.babec.org/files/PCR_2012/D1S80_PCR_Student_Guide_2012.pdf
My student versions of how to use Excel and other documents pertaining to this lab may be found:
- Graphing a standard curve
- Graphing absolute zero
- Making a solubility graph for KNO3
- NCBI database
Graphing a standard curve: How to graph the results of a d1s80 PCR lab and calculate the size of your bands.
One of my favorite lessons in biotechnology was the d1s80 PCR lab because the allele frequencies are known for its 40+ alleles. Students ran PCR reactions on their own DNA in our class. One of the labs looked at D1s80, a specific site in our genome. There are a few reasons we do this lab. First, it offers more experience with doing PCR labs. Second, the difference in allele sizes is only 16 base pairs. Being so small, we had to use polyacrylamide vertical gel boxes just to be able to see the bands. We ran a 50 bp or 100 bp ladder so that we could take the measurement of the distance migrated for bands of known length. DNA moves logarithmically. When we know the size of the band, we can calculate the log of its size. This is when Excel comes in to play. I tried to do this with Google spreadsheets, but it could not handle either doing the log or the antilog. I just know that Google spreadsheets could not handle this level of analysis in 2009. We set up the log of the base pair length for the bands on the y axis and distance migrated in centimeters on the x axis. After we do a linear regression (line of best fit), Excel will give us an equation. We take the distance our bands migrated and plug it in to the equation for the value of x. Solving the equation gives us the log of the bp of our band. We take the anti-log and we know how big our bands are for d1S80. This is the main part of the lab that deals with spreadsheets. We would then go on and calculate the genotype frequency using data on the allele frequencies of all known band lengths, but I'm not sure elaborating on that falls into this lesson. You should be able to access the important parts of the lab from links I put in this explanation so if you wanted to create a hypothetical lab for your students, you could. The third reason for doing this lab, in case it is not obvious, is the exposure I could give students with not only how to use Excel, but let them get a taste for how powerful it can be. While doing a linear regression is not so amazing, I think it is for high school students. They'll probably have to do at least one in college. Having a clue what one is and how to do one before they get to college, can only prepare them for the competition they will face in their classes.
To see the student version of the curricula, go here: http://www.babec.org/files/PCR_2012/D1S80_PCR_Student_Guide_2012.pdf
My student versions of how to use Excel and other documents pertaining to this lab may be found:
- D1s80 Excel graph
- Instructions on how to do the spreadsheet in Excel and the graphing- this is a tutorial I wrote for them that details how to do the d1s80 work. This was written in 2008, and because my version was slightly different than the one in our school's computer lab, there were problems. If you decide to use my guide, please be aware that it is for an older version of Excel since this was created in 2008.
- Sizes of the alleles for d1s80 (pdf)
- Allele frequencies for d1s80 (pdf)
- Data points on the graph
- A linear regression based on the points, including an equation

Graphing Absolute Zero
One of the best things about teaching chemistry is that on the scale we do it in high school, the relationship between pressure, volume, and temperature is linear and can be manipulated with algebra. In class we do a demonstration that involves students reading the values on the equipment so it is not fully teacher centric. I bought the apparatus at Fisher so I am using a screenshot of their object. I could not find a picture of it with a creative commons reuse label. The silver bulb, as you can see in the image, gets immersed in a container. We set up three or more temperatures so that we can graph our values. Two points may make a line, but you need at least three to have a trend.
So we measure the pressure and temperature of boiling water. The apparatus measures pressure- there is a valve that lets us make a constant volume so we can ignore the volume variable. A thermometer is used to measure the temperature. We measure room temperature and pressure. Finally we measure the pressure and temperature of the gas trapped in the apparatus at a cold temperature. You can minimally use ice cubes to get the cold temp. If you have access to dry ice, doing an alcohol and dry ice bath would be excellent.
Extrapolating the line to where it crosses the y axis should give you a value of – 273 Celsius. This value was reconfigured on the Kelvin scale and it is called 0 Kelvin. The “degree” label is not use with Kelvin degrees. I don’t know why. Our three or four data points let us calculate the value of Absolute Zero, 0 Kelvin. It is literally impossible to measure something at zero (0) K because all motion is stopped at 0 K. There is no way to actually have matter exist at 0 K, and be useful. We use math to calculate the value for absolute zero.
This is one of my favorite labs to do because it shows students how we can use evidence and extrapolate it to theoretical values. Reflecting on this assignment, it may not be the best one to demonstrate how to use spreadsheets and databases because students can do the graphing by hand. Also, there are not so many values that we’d need a spreadsheet to keep track of them. We could have students keep track of the data in a Google spreadsheet. I don’t think Google lets you do a graph of your data yet so you’d still be stuck with Excel or a similar program if you wanted to graph directly from the spreadsheet.
If you need values for a data table, which you can put into a spread sheet, this software will generate values for you: http://www.molecularsoft.com/gaslaws.htm In a way, this software functions as a database because you give it a value, and it determines the new temperature. Since this type of calculation should be easy to do, I made a Google spreadsheet and put in the formula for temperature 2. As long as the units for pressure are the same for each pressure value, then we can calculate the second temperature. I modelled this off of the Molecularsoft software. By the time this project goes live, I may figure out how to go from Fahrenheit to Celsius to Kelvin, and calculate Pressure 2 because that is usually how I see Gay-Lussac’s law happen. Usually the temperature changes, not the pressure.
My spreadsheet for Gay – Lussac’s law.
One of the best things about teaching chemistry is that on the scale we do it in high school, the relationship between pressure, volume, and temperature is linear and can be manipulated with algebra. In class we do a demonstration that involves students reading the values on the equipment so it is not fully teacher centric. I bought the apparatus at Fisher so I am using a screenshot of their object. I could not find a picture of it with a creative commons reuse label. The silver bulb, as you can see in the image, gets immersed in a container. We set up three or more temperatures so that we can graph our values. Two points may make a line, but you need at least three to have a trend.
So we measure the pressure and temperature of boiling water. The apparatus measures pressure- there is a valve that lets us make a constant volume so we can ignore the volume variable. A thermometer is used to measure the temperature. We measure room temperature and pressure. Finally we measure the pressure and temperature of the gas trapped in the apparatus at a cold temperature. You can minimally use ice cubes to get the cold temp. If you have access to dry ice, doing an alcohol and dry ice bath would be excellent.
Extrapolating the line to where it crosses the y axis should give you a value of – 273 Celsius. This value was reconfigured on the Kelvin scale and it is called 0 Kelvin. The “degree” label is not use with Kelvin degrees. I don’t know why. Our three or four data points let us calculate the value of Absolute Zero, 0 Kelvin. It is literally impossible to measure something at zero (0) K because all motion is stopped at 0 K. There is no way to actually have matter exist at 0 K, and be useful. We use math to calculate the value for absolute zero.
This is one of my favorite labs to do because it shows students how we can use evidence and extrapolate it to theoretical values. Reflecting on this assignment, it may not be the best one to demonstrate how to use spreadsheets and databases because students can do the graphing by hand. Also, there are not so many values that we’d need a spreadsheet to keep track of them. We could have students keep track of the data in a Google spreadsheet. I don’t think Google lets you do a graph of your data yet so you’d still be stuck with Excel or a similar program if you wanted to graph directly from the spreadsheet.
If you need values for a data table, which you can put into a spread sheet, this software will generate values for you: http://www.molecularsoft.com/gaslaws.htm In a way, this software functions as a database because you give it a value, and it determines the new temperature. Since this type of calculation should be easy to do, I made a Google spreadsheet and put in the formula for temperature 2. As long as the units for pressure are the same for each pressure value, then we can calculate the second temperature. I modelled this off of the Molecularsoft software. By the time this project goes live, I may figure out how to go from Fahrenheit to Celsius to Kelvin, and calculate Pressure 2 because that is usually how I see Gay-Lussac’s law happen. Usually the temperature changes, not the pressure.
My spreadsheet for Gay – Lussac’s law.
|
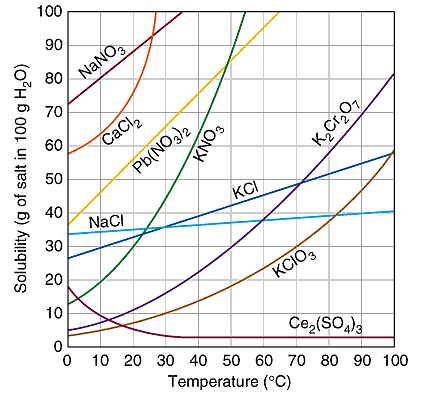
KNO3 graph
A lab we do as a class is the solubility graph of KNO3. I did not tell the students it was potassium nitrate, though. As a class we gathered data and graphed it. The solubility curve for KNO3 is so characteristic that it should be easily identified by comparing it to known solubility curves. This particular lab is here because it lends itself well to a spreadsheet. I did this lab in around the year 2000, so having a common Google doc was not an option. If I were to do this today, depending on what resources were available at my school, I would like to have students input their data into a Google spreadsheet. I would want them to program calculations into the cells. Like I keep saying, I don’t know of an online graphing program so they may still have to do the graphing by hand. a copy of the lab done in class image from: http://www.sciencegeek.net/Chemistry/taters/graphics/solubility.gif |

NCBI database
NCBI is the National Center for Biotechnology Information. If you have been reading my posts or projects, you may be getting used to the idea that I am thrilled to be as old as I am because it means I have been able to witness and experience changes the Internet has allowed us to have. I did my first Masters degree in 1991-1994. It was a PhD program which I quit and on a whim decided to become a science teacher. Since it was a research based Masters, I did research for two and a half years. One thing I did was sequencing. The virus I was working on was a RNA virus and RNA sequencing had not been perfected yet so I had to first make a DNA copy of the region I was sequencing. I bring this up because before the Internet allowed for “free” research tools, we used to have to remotely dial into specific servers to have our gene sequence analyzed. Doing BLAST searches has been around for a long time. Twenty years ago, however, you had to wait your turn on the server in Wisconsin (I believe that is where it was). Just like it could take 3 days for me to get the results from a sequencing gel due to having to radioactively label a nucleotide, and the autorads did not always work the first time, I had to be patient with getting sequence information about my virus. Life was much slower twenty three years ago.
The NCBI database is one of my favorite databases because you can input any sequence and analyze it immediately. Your result comes back in seconds, although sometimes you may have to wait a minute or two if it is a long sequence. I actually had students sequence a small region of their mitochondrial DNA and for the few students who it worked with, we did a small comparison of sequences in the NCBI database. I had them compare to other species to show how there are similarities among all species, but they should be happy sometimes when they get back a BLAST search and nothing matched up. I’ve included in this area some of the handouts I used with the students. I also uploaded some of the data so you can see how we have to respect students’ privacy. I have no clue what name goes with these numbers, but somehow, somewhere students knew which sequence was theirs in the sequence database. Sequencing gels used to be run at Cold Spring Harbor Laboratories (CSHL). It has been six years since I’ve done the mtDNA lab with students so I have no clue what they do now. One year we went to the local university and ran our products on their sequencing gels, which then transferred the information to the CSHL server. Another year I drove a couple students to what was at that time Applied Biosystems, and we ran our results on their sequencing gel. Applied Biosystems and Cold Spring Harbor Laboratories were huge supporters of this lab, and I will always be grateful for the opportunities they gave to my students.
Links and resources:
Image in this section is from the NCBI website: http://www.ncbi.nlm.nih.gov/About/images/splash_r2_c3.gif
NCBI is the National Center for Biotechnology Information. If you have been reading my posts or projects, you may be getting used to the idea that I am thrilled to be as old as I am because it means I have been able to witness and experience changes the Internet has allowed us to have. I did my first Masters degree in 1991-1994. It was a PhD program which I quit and on a whim decided to become a science teacher. Since it was a research based Masters, I did research for two and a half years. One thing I did was sequencing. The virus I was working on was a RNA virus and RNA sequencing had not been perfected yet so I had to first make a DNA copy of the region I was sequencing. I bring this up because before the Internet allowed for “free” research tools, we used to have to remotely dial into specific servers to have our gene sequence analyzed. Doing BLAST searches has been around for a long time. Twenty years ago, however, you had to wait your turn on the server in Wisconsin (I believe that is where it was). Just like it could take 3 days for me to get the results from a sequencing gel due to having to radioactively label a nucleotide, and the autorads did not always work the first time, I had to be patient with getting sequence information about my virus. Life was much slower twenty three years ago.
The NCBI database is one of my favorite databases because you can input any sequence and analyze it immediately. Your result comes back in seconds, although sometimes you may have to wait a minute or two if it is a long sequence. I actually had students sequence a small region of their mitochondrial DNA and for the few students who it worked with, we did a small comparison of sequences in the NCBI database. I had them compare to other species to show how there are similarities among all species, but they should be happy sometimes when they get back a BLAST search and nothing matched up. I’ve included in this area some of the handouts I used with the students. I also uploaded some of the data so you can see how we have to respect students’ privacy. I have no clue what name goes with these numbers, but somehow, somewhere students knew which sequence was theirs in the sequence database. Sequencing gels used to be run at Cold Spring Harbor Laboratories (CSHL). It has been six years since I’ve done the mtDNA lab with students so I have no clue what they do now. One year we went to the local university and ran our products on their sequencing gels, which then transferred the information to the CSHL server. Another year I drove a couple students to what was at that time Applied Biosystems, and we ran our results on their sequencing gel. Applied Biosystems and Cold Spring Harbor Laboratories were huge supporters of this lab, and I will always be grateful for the opportunities they gave to my students.
Links and resources:
- NCBI website, BLAST area
- student manual for mtDNA lab from BABEC
- data sheet from lab with kids
- another data sheet
- lesson on comparing their sequences to other animals
Image in this section is from the NCBI website: http://www.ncbi.nlm.nih.gov/About/images/splash_r2_c3.gif
OOPS!
After finishing this assignment and realizing I may have done it wrong, I want to put a few database connections here:
Some databases that were not mentioned already in this lesson or the accompanying blog include:
After finishing this assignment and realizing I may have done it wrong, I want to put a few database connections here:
Some databases that were not mentioned already in this lesson or the accompanying blog include:

CHEMnetBase has links to several chemical databases online. One that may be familiar to those who have taken college chemistry is the Handbook of Chemistry and Physics. They also have databases that have natural product inventories or even drugs. They also have dictionaries of food substances and of organic compounds.
Protein Data Bank, Molecule of the Month- I was going to use this database in one of the extra credit opportunities because I have students pick a molecule from this database and present it to the class. The Protein Data Bank (PDB) is also well known for its images of pretty much any protein molecule. One of the more fun areas to visit there are the animations.
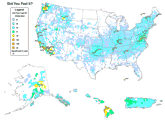
USGS- earthquake data and maps
This may not seem very significant, but there are now more reasons to be interested in small earthquakes. They are caused by fracking. You can have students map where earthquakes are happening in the US and explain why there are so many "new" earthquake sites. I happen to live in an area where the old fashioned earthquakes happened, not the ones due to water being forced underground. My favorite page after a local quake is Did You Feel It? because I am often curious how far it was felt and if others in my area thought it was as strong as I did.
When teaching earth science or geography, this website can be useful to identify where in the world earthquakes are likely to occur. Many years ago I had my students do a map of earthquake locations so they could identify the "ring of fire" and why it would be given that name.
This may not seem very significant, but there are now more reasons to be interested in small earthquakes. They are caused by fracking. You can have students map where earthquakes are happening in the US and explain why there are so many "new" earthquake sites. I happen to live in an area where the old fashioned earthquakes happened, not the ones due to water being forced underground. My favorite page after a local quake is Did You Feel It? because I am often curious how far it was felt and if others in my area thought it was as strong as I did.
When teaching earth science or geography, this website can be useful to identify where in the world earthquakes are likely to occur. Many years ago I had my students do a map of earthquake locations so they could identify the "ring of fire" and why it would be given that name.
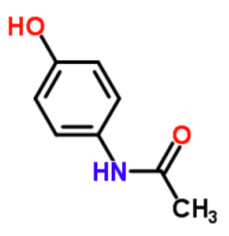 Paracetamol, a more scientific name than Tylenol, which is what I put in the database.
Paracetamol, a more scientific name than Tylenol, which is what I put in the database.
In case it is not obvious by now, I love the Royal Society of Chemistry. I promoted their interactive periodic table in another assignment. For the database assignment, I want to promote their ChemSpider. ChemSpider lets you put in a name of a chemical and it gives you back the molecular structure, other common or scientific names for the molecule, some of its reported physical and chemical properties, and even its spectral analysis. If I was in college now, I am not sure I'd be using the Handbook of Chemistry and Physics as often as I'd be using ChemSpider. It even has links to molecular structures and loads jmol (software that lets you manipulate the structures) easily. Not only will you get data about your molecule, but you can rotate it in 3D.

Part 2: A lesson using a spreadsheet.
Gallagher et al. (2011) remarked in their article that “computer science has become ubiquitous in many areas of biological research.” They go on to comment on how they are the first ones to do the types of lessons they are writing about. Looking at their curriculum, I think I can see why they say they are the first ones to do this type of lesson. They start off by teaching what an algorithm is. At first I thought I may adapt their lesson plan to be the one I present for this assignment, but now I’m not so sure I want to take the direction they are taking. They do, however, have Python code available to show students how an algorithm can work. I could not get it to run; however, I may not be using the correct compiler. Their second lesson focuses on doing a BLAST search, which is what I originally thought I would focus on for this lesson. They chose to use sickle cell anemia as the disease to research. They also provide students with human sequence data and rat sequence data, with the idea the students will do a BLAST search with each. After doing an alignment with the rat and human DNA, they have students do a full BLAST search with the rat DNA. This is the type of search I want to do, if for no other reason to try to bring some freedom for students to choose the organism they want to research. Their third lesson teaches students to create a phylogenetic tree. OK, now I’m really impressed. They provide a dozen sequences that the students use in comparison relationships to figure out who is more closely related in terms of evolutionary differences in sequence. The idea is that the more similar the sequences, the closer the two specimens are related genetically. I love this phylogenetic tree idea.
Upon doing more research for curriculum that has already been written, I found that there is actually quite a lot available for teachers who want to do bioinformatics with their students. The following websites contain tidbits of information I may utilize when crafting the spreadsheet project. Form and Lewitter (2011) wrote an article advising how to teach bioinformatics at the high school level. I will be using their suggestions as I create my lesson for this assignment. Gasstationwithoutpumps has a wordpress blog, but does not give a real name so I can’t figure out how to properly acknowledge his work. His post provides many resources that are useful for those who intend to use a bioinformatics lesson. The Northwest Association for Biomedical Research had a NSF grant to develop bioinformatics lessons. Their curriculum is thorough with lots of support material for how to understand what is involved with genetic testing.
After perusing what is already available, and thinking about the challenge teachers have had with making the information not be too overwhelming, yet still allow for student choice in what they explore, I have decided to have the spreadsheet lesson be one where the students mentally dissect GenBank entries. I chose GenBank entries because they have identifying information that the students would use if they actually do a BLAST alignment. The GenBank page lists the organism's scientific and common name, if it has one. It tells where it is located in the genome of the organism, the size of the fragment recorded for this entry, the type of sequence (DNA, protein, RNA), the GenBank category, and the date it was uploaded. In addition to this demographic type of information, it will have whatever sequence information is available to compliment what was uploaded. After using the spreadsheet, students should be aware that all organisms have DNA. They should be more comfortable with understanding there is a database of sequences they can look up. A follow-up lesson to this one would have students pick two of the organisms that they put in their database and do an alignment with them. At this point, I am happy if this gets students curious about pursuing this in college or as a career. We do not have enough data scientists to mine all sequence data that has been created. We need people on both sides- those who want to sequence genomes, and those who want to make sense of the sequences. I want students to know this is something they can do if they want to.
The lesson:
1. Go to: http://www.ncbi.nlm.nih.gov/
2. Click on DNA & RNA
3. Go to the database list.
4. Find a link to GenBank in their database list. (http://www.ncbi.nlm.nih.gov/genbank/)
5. Read about GenBank.
6. Open the annotated sample GenBank record for Saccharomyces cerevisiae (common yeast)
7. Look at the data in that page.
8. Identify what the following items mean or why they are a part of the record:
a. Locus
b. Definition
c. Accession
d. Version
e. Source Organism
f. Reference with Authors, title, journal, and pubmed- why is this information here?
g. In CDS, they have a translation section. There are a bunch of letters there. What do the letters represent?
h. In the Origin section there are another set of letters. What do these letters represent?
9. Based on the information that is available in a query, we will be making a database of information you can get from these pages.
10. The most important information is in the Locus line. The information from the Saccharomyces cerevisiae has been done for you.
11. Your assignment is to find GenBank entries for five of the GenBank divisions.:
a. PRI - primate sequences
b. ROD - rodent sequences
c. MAM - other mammalian sequences (not primate or rodent)
d. VRT - other vertebrate sequences (not primate or rodent)
e. INV - invertebrate sequences (like insects, crustaceans, or shellfish)
f. PLN - plant, fungal, and algal sequences
g. BCT - bacterial sequences
h. VRL - viral sequences
i. PHG - bacteriophage sequences
j. PAT - patent sequences (this may come up in a search so I am listing it as an option)
k. CON stands for contigs- these are short fragments of areas that have been sequenced many times. Their actual sequence has not been resolved yet. Only use a CON category if you absolutely can't find an organism in one of the more common types of divisions, like plants, animals, bacteria, or viruses.
12. If you can't magically think of an organism to fit in a category, you can Google the category name like: sequenced primate genomes. A long list of sequenced genomes should come up. Change the word, "primate" to fit the category you are trying to find.
13. From there, choose a link that will lead you to a name of an organism you can use.
14. Enter this name in the GenBank search box at the top of the page. I usually leave it on "Nucleotide" to do the searches. I have no clue what you will get back if you use a different category.
15. Fill in the table with the information with the information that is requested in the spreadsheet columns. Be sure to include the real name of the division in addition to its three letter abbreviation. The embedded form is difficult to use so I recommend using the one that opens up with the link in this part.
Gallagher et al. (2011) remarked in their article that “computer science has become ubiquitous in many areas of biological research.” They go on to comment on how they are the first ones to do the types of lessons they are writing about. Looking at their curriculum, I think I can see why they say they are the first ones to do this type of lesson. They start off by teaching what an algorithm is. At first I thought I may adapt their lesson plan to be the one I present for this assignment, but now I’m not so sure I want to take the direction they are taking. They do, however, have Python code available to show students how an algorithm can work. I could not get it to run; however, I may not be using the correct compiler. Their second lesson focuses on doing a BLAST search, which is what I originally thought I would focus on for this lesson. They chose to use sickle cell anemia as the disease to research. They also provide students with human sequence data and rat sequence data, with the idea the students will do a BLAST search with each. After doing an alignment with the rat and human DNA, they have students do a full BLAST search with the rat DNA. This is the type of search I want to do, if for no other reason to try to bring some freedom for students to choose the organism they want to research. Their third lesson teaches students to create a phylogenetic tree. OK, now I’m really impressed. They provide a dozen sequences that the students use in comparison relationships to figure out who is more closely related in terms of evolutionary differences in sequence. The idea is that the more similar the sequences, the closer the two specimens are related genetically. I love this phylogenetic tree idea.
Upon doing more research for curriculum that has already been written, I found that there is actually quite a lot available for teachers who want to do bioinformatics with their students. The following websites contain tidbits of information I may utilize when crafting the spreadsheet project. Form and Lewitter (2011) wrote an article advising how to teach bioinformatics at the high school level. I will be using their suggestions as I create my lesson for this assignment. Gasstationwithoutpumps has a wordpress blog, but does not give a real name so I can’t figure out how to properly acknowledge his work. His post provides many resources that are useful for those who intend to use a bioinformatics lesson. The Northwest Association for Biomedical Research had a NSF grant to develop bioinformatics lessons. Their curriculum is thorough with lots of support material for how to understand what is involved with genetic testing.
After perusing what is already available, and thinking about the challenge teachers have had with making the information not be too overwhelming, yet still allow for student choice in what they explore, I have decided to have the spreadsheet lesson be one where the students mentally dissect GenBank entries. I chose GenBank entries because they have identifying information that the students would use if they actually do a BLAST alignment. The GenBank page lists the organism's scientific and common name, if it has one. It tells where it is located in the genome of the organism, the size of the fragment recorded for this entry, the type of sequence (DNA, protein, RNA), the GenBank category, and the date it was uploaded. In addition to this demographic type of information, it will have whatever sequence information is available to compliment what was uploaded. After using the spreadsheet, students should be aware that all organisms have DNA. They should be more comfortable with understanding there is a database of sequences they can look up. A follow-up lesson to this one would have students pick two of the organisms that they put in their database and do an alignment with them. At this point, I am happy if this gets students curious about pursuing this in college or as a career. We do not have enough data scientists to mine all sequence data that has been created. We need people on both sides- those who want to sequence genomes, and those who want to make sense of the sequences. I want students to know this is something they can do if they want to.
The lesson:
1. Go to: http://www.ncbi.nlm.nih.gov/
2. Click on DNA & RNA
3. Go to the database list.
4. Find a link to GenBank in their database list. (http://www.ncbi.nlm.nih.gov/genbank/)
5. Read about GenBank.
6. Open the annotated sample GenBank record for Saccharomyces cerevisiae (common yeast)
7. Look at the data in that page.
8. Identify what the following items mean or why they are a part of the record:
a. Locus
b. Definition
c. Accession
d. Version
e. Source Organism
f. Reference with Authors, title, journal, and pubmed- why is this information here?
g. In CDS, they have a translation section. There are a bunch of letters there. What do the letters represent?
h. In the Origin section there are another set of letters. What do these letters represent?
9. Based on the information that is available in a query, we will be making a database of information you can get from these pages.
10. The most important information is in the Locus line. The information from the Saccharomyces cerevisiae has been done for you.
11. Your assignment is to find GenBank entries for five of the GenBank divisions.:
a. PRI - primate sequences
b. ROD - rodent sequences
c. MAM - other mammalian sequences (not primate or rodent)
d. VRT - other vertebrate sequences (not primate or rodent)
e. INV - invertebrate sequences (like insects, crustaceans, or shellfish)
f. PLN - plant, fungal, and algal sequences
g. BCT - bacterial sequences
h. VRL - viral sequences
i. PHG - bacteriophage sequences
j. PAT - patent sequences (this may come up in a search so I am listing it as an option)
k. CON stands for contigs- these are short fragments of areas that have been sequenced many times. Their actual sequence has not been resolved yet. Only use a CON category if you absolutely can't find an organism in one of the more common types of divisions, like plants, animals, bacteria, or viruses.
12. If you can't magically think of an organism to fit in a category, you can Google the category name like: sequenced primate genomes. A long list of sequenced genomes should come up. Change the word, "primate" to fit the category you are trying to find.
13. From there, choose a link that will lead you to a name of an organism you can use.
14. Enter this name in the GenBank search box at the top of the page. I usually leave it on "Nucleotide" to do the searches. I have no clue what you will get back if you use a different category.
15. Fill in the table with the information with the information that is requested in the spreadsheet columns. Be sure to include the real name of the division in addition to its three letter abbreviation. The embedded form is difficult to use so I recommend using the one that opens up with the link in this part.
- Scientific name of organism
- Common name of organism, if given. If not, the description from the definition.
- Locus identifying number
- Size of fragment being recorded. If you used the nucleotide bank with the search box, this value is most likely the number of bases in the sequence
- Type of sequence (DNA, cDNA, mRNA). Please indicate genomic or mitochondrial, if possible.
- GenBank category (the one listed in the a-k categories mentioned above), (3 letters and description)
- Date sequence was uploaded
Resources:
Form, D., & Lewitter, F. (2011). Ten Simple Rules for Teaching Bioinformatics at the High School Level. PLoS Computational Biology, 7(10), e1002243. doi:10.1371/journal.pcbi.1002243
Gallagher, S. R., Coon, W., Donley, K., Scott, A., & Goldberg, D. S. (2011). A First Attempt to Bring Computational Biology into Advanced High School Biology Classrooms. PLoS Computational Biology, 7(10), e1002244. doi:10.1371/journal.pcbi.1002244
Introductory bioinformatics: Genetic testing. (2013, April 22). Retrieved February 20, 2015, from https://www.nwabr.org/teacher-center/introductory-bioinformatics-genetic-testing#lessons
PLoS computational biology: Bioinformatics: Starting Early. (2012, February 25). Retrieved February 20, 2015, from https://gasstationwithoutpumps.wordpress.com/2012/02/25/plos-computational-biology-bioinformatics-starting-early/
Form, D., & Lewitter, F. (2011). Ten Simple Rules for Teaching Bioinformatics at the High School Level. PLoS Computational Biology, 7(10), e1002243. doi:10.1371/journal.pcbi.1002243
Gallagher, S. R., Coon, W., Donley, K., Scott, A., & Goldberg, D. S. (2011). A First Attempt to Bring Computational Biology into Advanced High School Biology Classrooms. PLoS Computational Biology, 7(10), e1002244. doi:10.1371/journal.pcbi.1002244
Introductory bioinformatics: Genetic testing. (2013, April 22). Retrieved February 20, 2015, from https://www.nwabr.org/teacher-center/introductory-bioinformatics-genetic-testing#lessons
PLoS computational biology: Bioinformatics: Starting Early. (2012, February 25). Retrieved February 20, 2015, from https://gasstationwithoutpumps.wordpress.com/2012/02/25/plos-computational-biology-bioinformatics-starting-early/